Learning Objectives¶
Import the Pandas library and understand when to use it over NumPy.
Use Pandas to load a CSV data set.
Get basic information about a Pandas DataFrame using
info(),describe(),columns, andT.Set a column as the row index using
DataFrame.set_index.Select individual values from a DataFrame using
iloc(by position) andloc(by label).Select a subset of both rows and columns from a DataFrame in a single operation.
Select a subset of a DataFrame by a single Boolean criterion (masking).
Make quick plots directly from a DataFrame using
DataFrame.plot().Understand Python dictionaries and how to iterate over their keys.
Create a DataFrame from a dictionary.
Combine multiple DataFrames using
pd.concat.Save a DataFrame to a CSV file using
DataFrame.to_csv.
Using the Pandas library to do statistics on tabular data¶
Pandas is a an open source library providing high-performance, easy-to-use data structures and data analysis tools. Pandas is particularly suited to the analysis of tabular data, i.e. data that can can go into a table. In other words, if you can imagine the data in an Excel spreadsheet, then Pandas is the tool for the job.
Pandas is a widely-used Python library for statistics, particularly on tabular data.
Borrows many features from R’s dataframes.
A 2-dimensional table whose columns have names and potentially have different data types.
Load it with import pandas as pd. The alias pd is commonly used for Pandas.
Read a Comma Separated Values (CSV) data file with pd.read_csv.
Argument is the name of the file to be read.
Assign result to a variable to store the data that was read.
In this lecture, we will go over the basic capabilities of Pandas. It is a very deep library, and you will need to dig into the documentation for more advanced usage.
import numpy as np
import pandas as pdReading data into a pandas dataframe¶
Lets use pandas to read one of our well-log data files
path = '/mnt/c/Ryan_Data/Teaching/work/classes/GPGN268/coursework-du/ds01-well-log/data/iodp-logging-data/EXP372/U1517A/372-U1517A_res-phase-nscope.csv'
df = pd.read_csv(path)---------------------------------------------------------------------------
ParserError Traceback (most recent call last)
Cell In[2], line 2
1 path = '/mnt/c/Ryan_Data/Teaching/work/classes/GPGN268/coursework-du/ds01-well-log/data/iodp-logging-data/EXP372/U1517A/372-U1517A_res-phase-nscope.csv'
----> 2 df = pd.read_csv(path)
File /mnt/c/Ryan_Data/Teaching/GPGN268/GPGN268-BOOK/.pixi/envs/default/lib/python3.14/site-packages/pandas/io/parsers/readers.py:873, in read_csv(filepath_or_buffer, sep, delimiter, header, names, index_col, usecols, dtype, engine, converters, true_values, false_values, skipinitialspace, skiprows, skipfooter, nrows, na_values, keep_default_na, na_filter, skip_blank_lines, parse_dates, date_format, dayfirst, cache_dates, iterator, chunksize, compression, thousands, decimal, lineterminator, quotechar, quoting, doublequote, escapechar, comment, encoding, encoding_errors, dialect, on_bad_lines, low_memory, memory_map, float_precision, storage_options, dtype_backend)
861 kwds_defaults = _refine_defaults_read(
862 dialect,
863 delimiter,
(...) 869 dtype_backend=dtype_backend,
870 )
871 kwds.update(kwds_defaults)
--> 873 return _read(filepath_or_buffer, kwds)
File /mnt/c/Ryan_Data/Teaching/GPGN268/GPGN268-BOOK/.pixi/envs/default/lib/python3.14/site-packages/pandas/io/parsers/readers.py:306, in _read(filepath_or_buffer, kwds)
303 return parser
305 with parser:
--> 306 return parser.read(nrows)
File /mnt/c/Ryan_Data/Teaching/GPGN268/GPGN268-BOOK/.pixi/envs/default/lib/python3.14/site-packages/pandas/io/parsers/readers.py:1947, in TextFileReader.read(self, nrows)
1940 nrows = validate_integer("nrows", nrows)
1941 try:
1942 # error: "ParserBase" has no attribute "read"
1943 (
1944 index,
1945 columns,
1946 col_dict,
-> 1947 ) = self._engine.read( # type: ignore[attr-defined]
1948 nrows
1949 )
1950 except Exception:
1951 self.close()
File /mnt/c/Ryan_Data/Teaching/GPGN268/GPGN268-BOOK/.pixi/envs/default/lib/python3.14/site-packages/pandas/io/parsers/c_parser_wrapper.py:215, in CParserWrapper.read(self, nrows)
213 try:
214 if self.low_memory:
--> 215 chunks = self._reader.read_low_memory(nrows)
216 # destructive to chunks
217 data = _concatenate_chunks(chunks, self.names)
File pandas/_libs/parsers.pyx:832, in pandas._libs.parsers.TextReader.read_low_memory()
File pandas/_libs/parsers.pyx:897, in pandas._libs.parsers.TextReader._read_rows()
File pandas/_libs/parsers.pyx:868, in pandas._libs.parsers.TextReader._tokenize_rows()
File pandas/_libs/parsers.pyx:885, in pandas._libs.parsers.TextReader._check_tokenize_status()
File pandas/_libs/parsers.pyx:2084, in pandas._libs.parsers.raise_parser_error()
ParserError: Error tokenizing data. C error: Expected 2 fields in line 5, saw 16
We see that the pandas printed things into a table but there is something weird about it. It seem like the first couple of lines have a different format. Let’s use the terminal to look at the data file. Open a new terminal window from Jupyter Lab and navigate to the data folder
cd ~/work/classes/GPGN268/coursework-villasboas/ds01-well-log/data/iodp-logging-data/EXP372/U1517A
cat 372-U1517A_cali-nscope.csv | headdf = pd.read_csv(path, skiprows=[0, 1, 2, 3, 5])dfNow, we read our data using pandas, but what type of object df is?
type(df)pandas.DataFrameBefore we were using numpy to read data into numpy arrays. With pandas, our data is stored as a pandas DataFrame.
pandas.DataFrame.set_index¶
It specifies a column’s values should be used as row headings
Now, we see that the row labels are currently numbers (1-1226), but we would really want the depths to be the row labels. We can use
df.set_index('DEPTH_LSF', inplace=True)
dfpandas.DataFrame.head() and tail()¶
Use pandas.DataFrame.head() to show the first few rows of the DataFrame
You can also specify the number of rows with
df.head(10)
df.head()You can also specify the number of rows with
df.tail(10)
df.tail()DataFrame.info()¶
to find out more about a dataframe.
df.info()<class 'pandas.DataFrame'>
Index: 1226 entries, 0.1348 to 186.8248
Data columns (total 15 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 P16B 1226 non-null float64
1 P16H 1226 non-null float64
2 P16L 1226 non-null float64
3 P22B 1226 non-null float64
4 P22H 1226 non-null float64
5 P22L 1226 non-null float64
6 P28B 1226 non-null float64
7 P28H 1226 non-null float64
8 P28L 1226 non-null float64
9 P34B 1226 non-null float64
10 P34H 1226 non-null float64
11 P34L 1226 non-null float64
12 P40B 1226 non-null float64
13 P40H 1226 non-null float64
14 P40L 1226 non-null float64
dtypes: float64(15)
memory usage: 153.2 KB
Here is the information given to us
This is a DataFrame
There are 1226 rows with values from 0.1348 to 186.8248
There are fourteen columns, each of which has two actual 64-bit floating point values.
We will talk later about null values, which are used to represent missing observations.
Uses 153.2 KB of memory.
DataFrame.columns¶
Is a variable that stores information about the dataframe’s columns
Note that this is data, not a method (It doesn’t have parentheses), so do not use () to try to call it.
df.columnsIndex(['P16B', 'P16H', 'P16L', 'P22B', 'P22H', 'P22L', 'P28B', 'P28H', 'P28L',
'P34B', 'P34H', 'P34L', 'P40B', 'P40H', 'P40L'],
dtype='str')DataFrame.T transpose a DataFrame¶
Sometimes want to treat columns as rows and vice versa.
Transpose (written .T) doesn’t copy the data, just changes the program’s view of it.
Like columns, it is a member variable.
df.TDataFrame.describe() give summary statistics about data¶
DataFrame.describe() gets the summary statistics of only the columns that have numerical data. All other columns are ignored, unless you use the argument include=‘all’.
df.describe()✅ Activity
Look at the table above and try to explain to one of your peers what exactly
df.describe()represents. Discuss the meaning of each summary statistics.
Note about Pandas DataFrames/Series¶
A DataFrame is a collection of Series; The DataFrame is the way Pandas represents a table, and Series is the data-structure Pandas use to represent a column.
Pandas is built on top of the Numpy library, which in practice means that most of the methods defined for Numpy Arrays apply to Pandas Series/DataFrames.
What makes Pandas so attractive is the powerful interface to access individual records of the table, proper handling of missing values, and relational-databases operations between DataFrames.
Selecting values¶
To access the values of the index column, we can do:
df.indexIndex([ 0.1348, 0.2872, 0.4396, 0.592, 0.7444, 0.8968, 1.0492,
1.2016, 1.354, 1.5064,
...
185.4532, 185.6056, 185.758, 185.9104, 186.0628, 186.2152, 186.3676,
186.52, 186.6724, 186.8248],
dtype='float64', name='DEPTH_LSF', length=1226)To access a value at the position [i,j] of a DataFrame, we have two options, depending on what is the meaning of i in use. Remember that a DataFrame provides an index as a way to identify the rows of the table; a row, then, has a position inside the table as well as a label, which uniquely identifies its entry in the DataFrame.
DataFrame.iloc[..., ...]¶
It selects values by their (entry) position
Can specify location by numerical index analogously to 2D version of character selection in strings.
# Selects the value in the first row and first column
df.iloc[0, 0]np.float64(0.6091384)DataFrame.loc[..., ...]¶
It selects values by their (entry) label.
Can specify location by row and/or column name/value.
# First value is the "index" value (i.e., depth)
# and second value is the row label
df.loc[186.5200, "P16B"]np.float64(1.283353)# All rows and P22H column
df.loc[:, "P22H"]DEPTH_LSF
0.1348 0.628182
0.2872 0.632340
0.4396 0.632921
0.5920 0.637017
0.7444 0.645322
...
186.2152 1.349495
186.3676 1.286108
186.5200 1.298547
186.6724 1.272658
186.8248 1.272654
Name: P22H, Length: 1226, dtype: float64Select multiple columns or rows using DataFrame.loc and a named slice¶
df.loc[0.2872:0.7444, "P16L":"P28B"]👀 In the above code, we discover that slicing using loc is inclusive at both ends, which differs from slicing using iloc, where slicing indicates everything up to but not including the final index.
Result of slicing can be used in further operations¶
All the statistical operators that work on entire dataframes work the same way on slices. For example, calculate max of a slice.
# Return the maximum for each column
df.loc[0.2872:0.7444, "P16L":"P28B"].max()P16L 0.682845
P22B 0.697899
P22H 0.645322
P22L 0.697899
P28B 0.709838
dtype: float64Use comparisons to select data based on value¶
Comparison is applied element by element.
Returns a similarly-shaped dataframe of True and False.
subset = df.loc[:, "P16B":"P16L"]
subsetsubset > 1Select values using a Boolean mask¶
A frame full of Booleans is sometimes called a mask because of how it can be used.
mask = subset > 1
subset[mask]This returnsthe value where the mask is true, and NaN (Not a Number) where it is false.
This is useful because NaNs are ignored by operations like max, min, average, etc.
subset[mask].describe()🤔 How does that compare with subset.describe()?¶
Make quick plots for data inspection with pandas¶
A pandas DataFrame has a method plot() which allows quick visualization. Note that for final, you should still use matplotlib and customize your graphs for optimal presentation, but plotting straight from pandas is a great way to take a first look at your data.
df.plot()<Axes: xlabel='DEPTH_LSF'>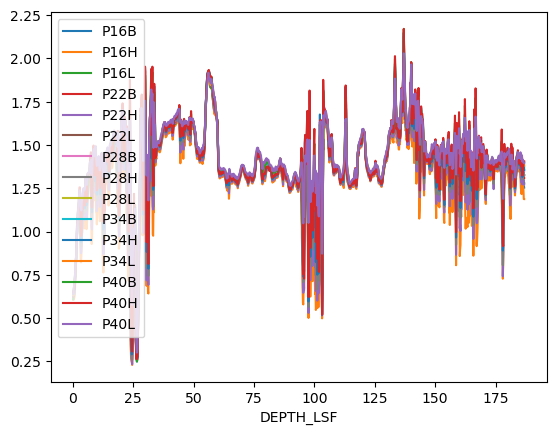
# You can also select only one variable to plot
df.loc[:, 'P22L'].plot(color='tomato', ls='--')<Axes: xlabel='DEPTH_LSF'>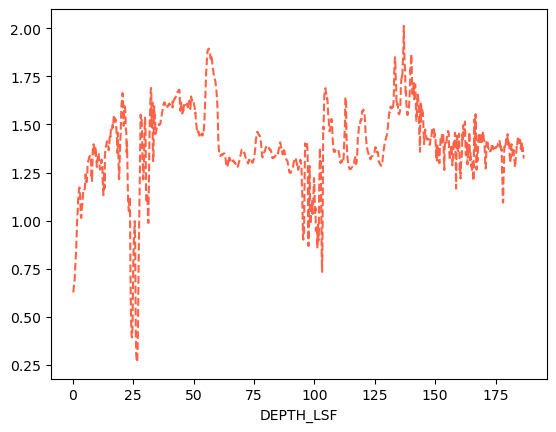
Change the label of the index column with index.name¶
df.index.name = 'Depth'dfDictionaries¶
So far we’ve worked with numpy arrays and pythin lists, but there is one important structure in Python that we havent discussed: dictionaries. This is an extremely useful data structure. It maps keys to values.
Dictionaries are unordered!
# Different ways to create disctionaries
# Using curly brackets-> key: value
band_facts = {'name': 'Coldplay', 'studio albums': 9}
# Usinf the function dict -> key=value
major_facts = dict(name='Geophysics', tracks=6)
print(type(band_facts))
print(band_facts)
print(type(major_facts))
print(major_facts)<class 'dict'>
{'name': 'Coldplay', 'studio albums': 9}
<class 'dict'>
{'name': 'Geophysics', 'tracks': 6}
To access values in a dictionary, you can’t use an index (because dictionaries are not ordered). You access values using their keys:
band_facts['name']'Coldplay'Square brackets [...] are Python for “get item” in many different contexts.
You can check for the presence of a key
# check if the dictionary major_facts has the key "tracks"
'tracks' in major_factsTrue# check if the dictionary major_facts has the key "students"
'students' in major_factsFalseIf you try to access a key that doesn’t exist in the dictionary, you will get an error:
# try to access a non-existant key
band_facts['grammys']---------------------------------------------------------------------------
KeyError Traceback (most recent call last)
Cell In[31], line 2
1 # try to access a non-existant key
----> 2 band_facts['grammys']
KeyError: 'grammys'You can add new keys to a dictonary:
band_facts['grammys'] = 7
band_facts{'name': 'Coldplay', 'studio albums': 9, 'grammys': 7}A very useful trick is to iterate over keys:
# iterate over keys
for k in band_facts:
print(k, band_facts[k])name Coldplay
studio albums 9
grammys 7
We were talking about pandas, so why we’ve suddenly change to dictionaries? Well it turns out dictionaries are fundamental to DataFrames. Let’s see how.
# first we create a dictionary
data = {'mass': [0.3e24, 4.87e24, 5.97e24], # kg
'diameter': [4879e3, 12_104e3, 12_756e3], # m
'rotation_period': [1407.6, np.nan, 23.9] # h
}
df = pd.DataFrame(data, index=['Mercury', 'Venus', 'Earth'])
dfPandas handles missing data very elegantly, keeping track of it through all calculations
df.info()<class 'pandas.DataFrame'>
Index: 3 entries, Mercury to Earth
Data columns (total 3 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 mass 3 non-null float64
1 diameter 3 non-null float64
2 rotation_period 2 non-null float64
dtypes: float64(3)
memory usage: 96.0+ bytes
As a recap, we can look at summary statistics using the describe method:
df.describe()We can get a single column, which will return a Pandas Series, using python’s getitem syntax on the DataFrame object.
df['mass']Mercury 3.000000e+23
Venus 4.870000e+24
Earth 5.970000e+24
Name: mass, dtype: float64# A Series is a one-dimensional pandas object
type(df['mass'])pandas.SeriesWe can also select a specific column using attribute (“dot”) syntax.
df.massMercury 3.000000e+23
Venus 4.870000e+24
Earth 5.970000e+24
Name: mass, dtype: float64We’ve already seen that we can index using the operations loc and iloc.
df# Select the row "Earth" and comlun "mass"
df.loc['Earth', 'mass']np.float64(5.97e+24)Merging, combining, and joining data¶
Pandas allow us to easly merge, combine, and manipulate DataFrames. Consider the dataframe below:
data = {'lead singer': ['Chris Martin', 'Freddie Mercury', 'Julian Casablancas'],
'albums': [9, 15, 6]
}
df1 = pd.DataFrame(data, index=['Coldplay', 'Queen', 'The Strokes'])
df1other_data = {'year created': [1997, 1970, 1998],
'grammys': [7, 0, 1]
}
df2 = pd.DataFrame(other_data, index=['Coldplay', 'Queen', 'The Strokes'])
df2Now, we would like to merge the two Dataframes keeping the index (rows) and merging along the columns (axis=1). First we create a list with the DataFrames we want to “stick together”
dfs = [df1, df2]Now we use the function concat, which stands for concatenate.
# We want to concatenate the DataFrames in the list "dfs" along the column axis (axis=1)
df_combined = pd.concat(dfs, axis=1)
df_combinedNow, let’s suppose we have some more Coldplay facts.
more_facts = {'number of members': 4,
'hometown': 'London'
}
df3 = pd.DataFrame(more_facts, index=['Coldplay'])
df3Now, we still want to aggregate that to our main DataFrame, but the row index now only has “Coldplay”. No problem! pandas will add the new fields to “Coldplay” and fill the rest with NaNs
df_all = pd.concat([df_combined, df3], axis=1)
df_allSaving your data¶
If you did quite a bit of manipulation on your raw data, it might be smart to save a processed version of it so you don’t have to repeat all the steps in your code everytime. It is straighforward to save a pandas DataFrame to a text file, such as csv. The syntax is:
df.to_csv('path/to/output/file_name.csv')For example, the line below saves our DataFrame on the Desktop with the name “band_facts.csv”.
df_all.to_csv('../data/band_facts.csv')Key Points¶
Use the Pandas library to do statistics on tabular data; load CSV files with
pd.read_csv.Use
DataFrame.set_indexto specify that a column’s values should be used as row headings.Use
DataFrame.infoto find out more about a dataframe.The
DataFrame.columnsvariable stores information about the dataframe’s columns.Use
DataFrame.Tto transpose a dataframe.Use
DataFrame.describeto get summary statistics about data.Use
DataFrame.ilocto select values by position; useDataFrame.locto select by label.locslicing is inclusive at both ends;ilocslicing excludes the final index.Use Boolean masks to filter data; masked-out values become NaN and are ignored in calculations like
max,min, andmean.Use
DataFrame.plot()for quick data visualization during exploration.Python dictionaries map keys to values and are the foundation for creating DataFrames manually.
Use
pd.concatto combine multiple DataFrames along rows (axis=0) or columns (axis=1); missing values are filled with NaN.Save a DataFrame to CSV with
DataFrame.to_csv('path/to/file.csv').